|
More specifically we omit the calculation of site-site interaction energies, and the calculation of desolvation and background interaction energies for a large number of pairs of titratable groups. We show that pKa values accurate enough for identifying the proton donor in an enzyme active site can be calculated by considering in detail only the active-site residues and their immediate electrostatic interaction partners, thus allowing for a large decrease in calculation time. Estimates of protein residue pKa values can be obtained by applying pKa calculation algorithms to protein X-ray structures. 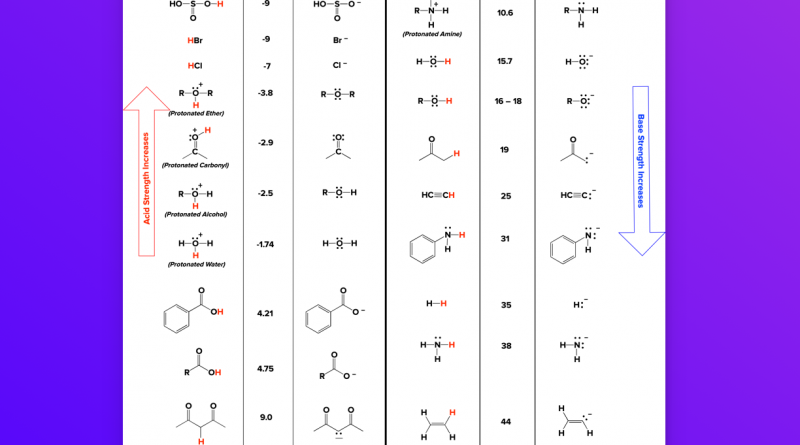 
Knowledge of these ionization constants (pKa values) often allows the researcher to identify the proton donor and the catalytic nucleophile in the reaction mechanism of the enzyme. The ionization properties of the active-site residues in enzymes are of considerable interest in the study of the catalytic mechanisms of enzymes.
0 Comments
Leave a Reply. |
 RSS Feed
RSS Feed
